PhenoMATRIX®
Advanced Artificial Intelligence for Microbiology
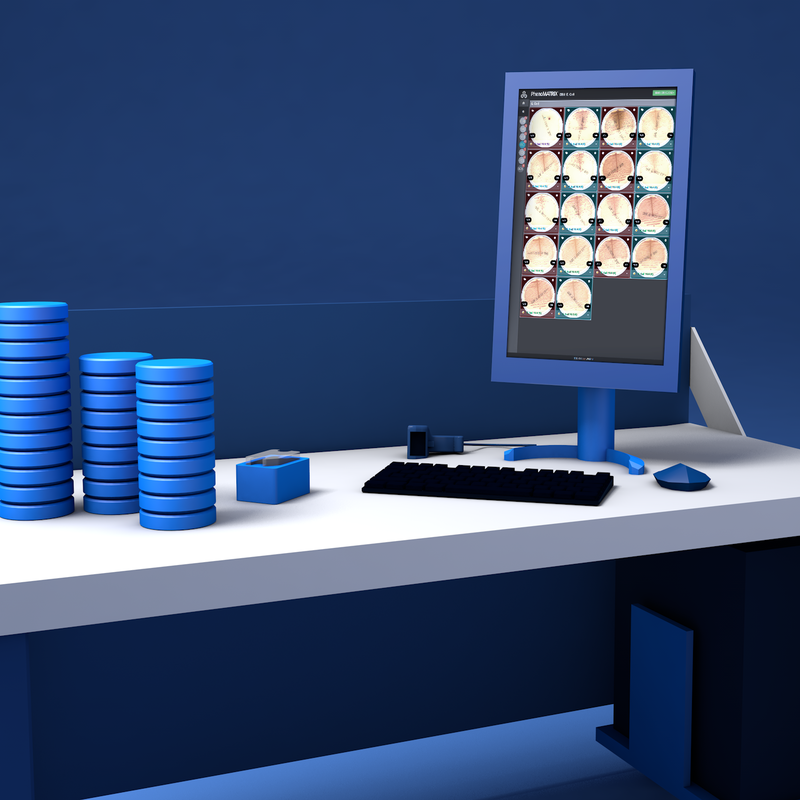

Product Overview
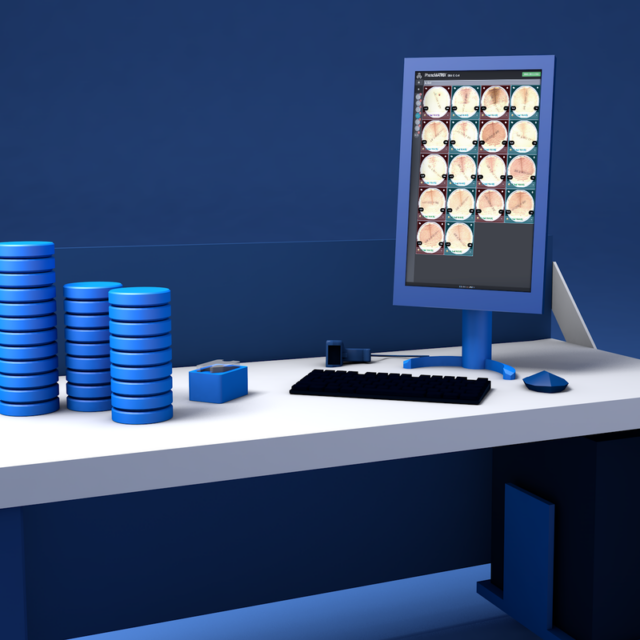
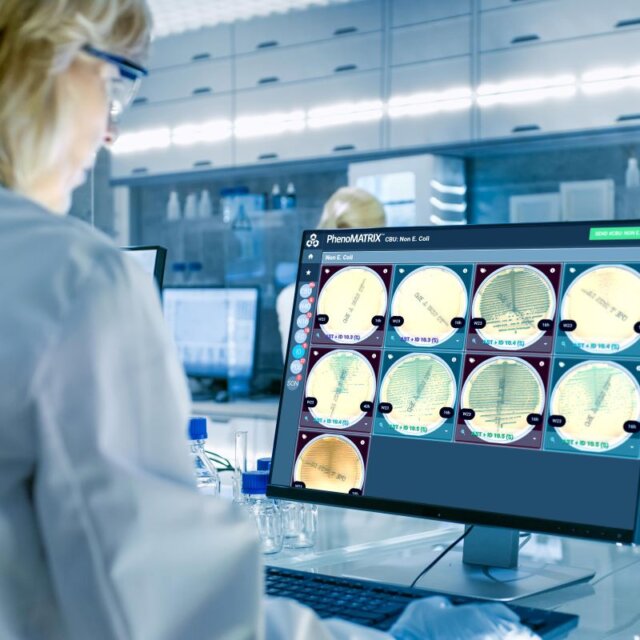
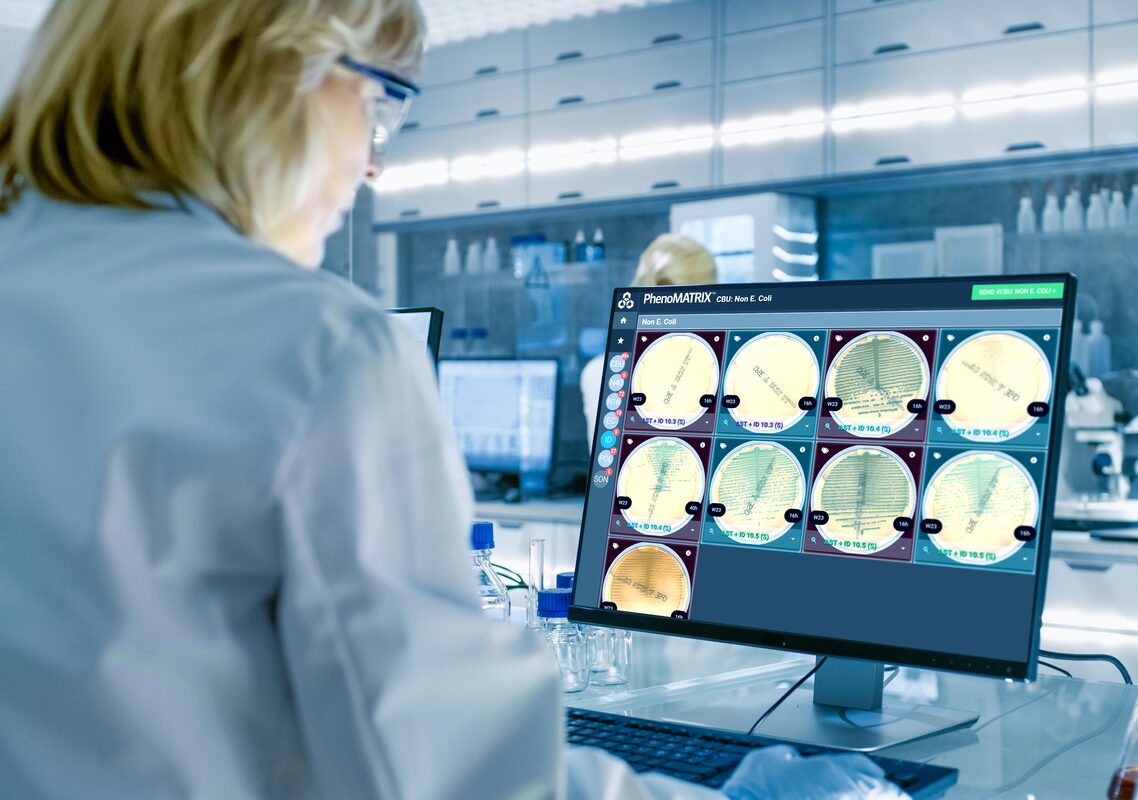
Unparalleled in the industry, PhenoMATRIX® is FDA 510(k)- cleared software that uses artificial intelligence, combined with patient information from the Laboratory Information System (LIS) and laboratory-defined expert rules, to automatically sort digital images of culture media plates and assist technologists with final assessment and result definition.
- Detects microbial growth, estimates semi-quantitative and qualitative colony counts, and differentiates isolates based on phenotypic colony characteristics
- Automatically sorts culture plate images according to laboratory-defined classification parameters
- Choose from multiple configuration options to meet your laboratory needs*
PhenoMATRIX® is FDA 510(k) cleared for its intended use. Certain additional configurations described below are labeled Research Use Only (RUO) in the United States.

Streamline Negative Plate Review
Help reduce time spent reviewing negative and insignificant cultures by automatically sorting images and allowing technologists to prioritize their time, for plates requiring further evaluation.1, 3

Increase Efficiency and Streamline Workflow
Automatically assign a suggested result to the culture plate images to support technologist to either accept or modify by reviewing the sorted folders and streamlined reporting to the LIS.1, 6

Improve Consistency and Reproducibility
Artificial Intelligence allows laboratory professionals to use their valuable time and expertise on critical thinking, analysis, and interpretation.2, 5, 6
Empower Laboratory Staff
Artificial Intelligence allows laboratory professionals to use their valuable time and expertise on critical thinking, analysis, and interpretation.1, 4
PhenoMATRIX® Receives FDA 510(k) Clearance
PhenoMATRIX® is now FDA 510(k) cleared (K251511) as a Class II Device, marking a significant milestone in digital microbiology innovation.
By combining image assessment with LIS-linked clinical context, PhenoMATRIX® supports standardized, efficient workflow while preserving technologist oversight.

Simplify Workflow with Intelligent Image Analysis
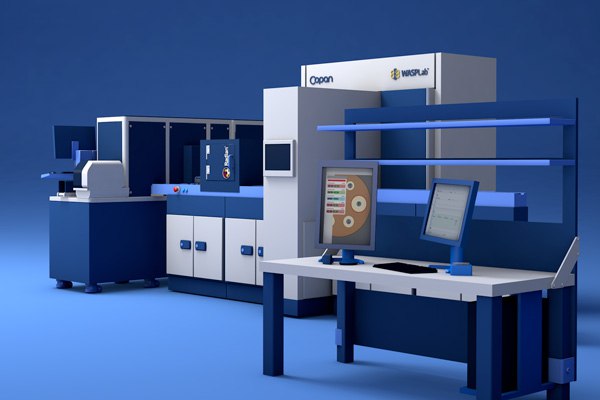
The PhenoMATRIX® AI suite supports automated image assessment and sorting of culture plate images within the WASPLab® digital workflow.
The system applies laboratory-defined expert rules and image analysis results to organize plates for efficient technologist review. Final assessment and result definition remain the responsibility of trained laboratory personnel.1, 6

Streamline Culture Analysis and Reporting
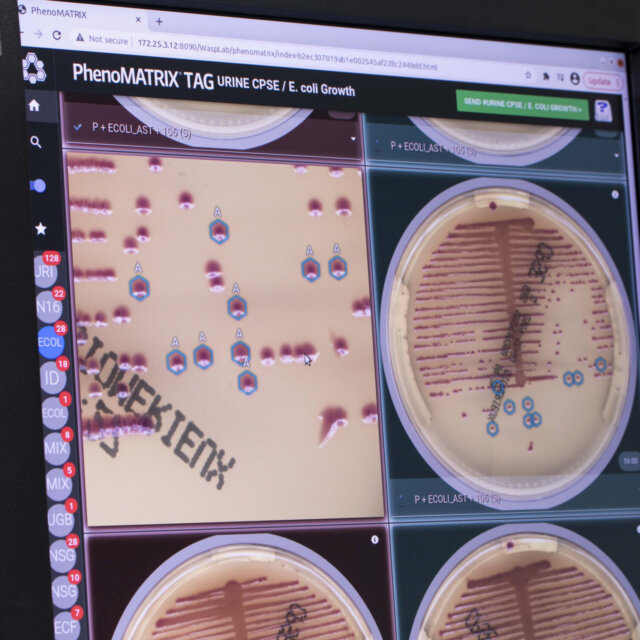
PhenoMATRIX® software analyzes high-resolution culture images and sorts them based on criteria including growth presence, growth semi-quantitative assessment, isolate quantitation, morphological interpretation, hemolysis patterns, and other laboratory-defined parameters.
This supports efficient review by organizing plates into laboratory-defined categories before technologist evaluation. Laboratory technologists remain responsible for reviewing all images and, following laboratory protocols, can efficiently report grouped negative and positive results directly to the LIS with a simple click. 3, 4-6
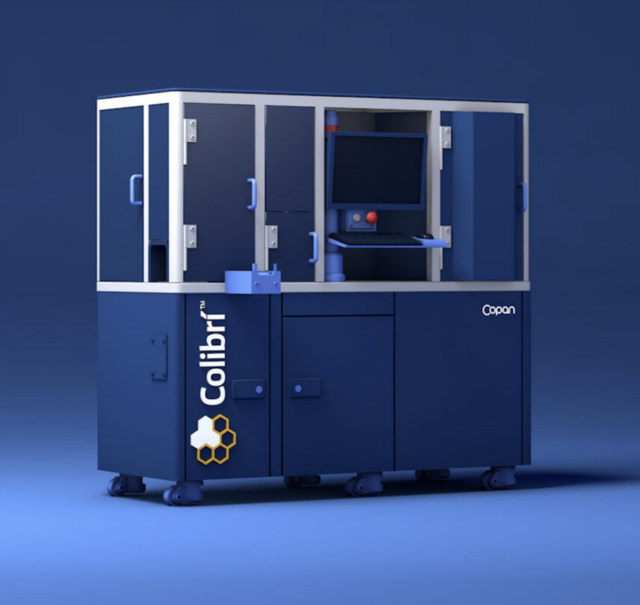
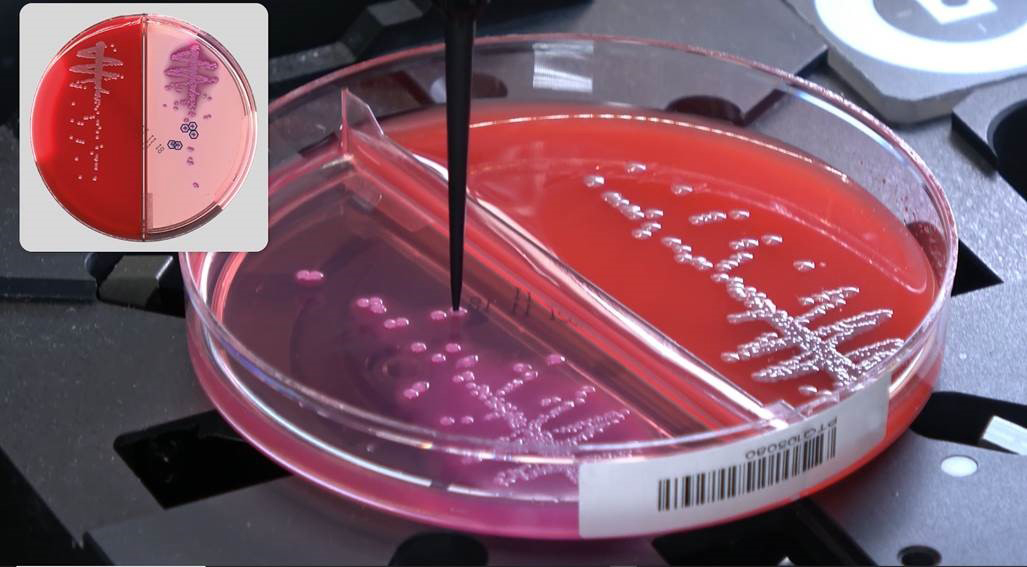
PhenoMATRIX® and Colibrí™
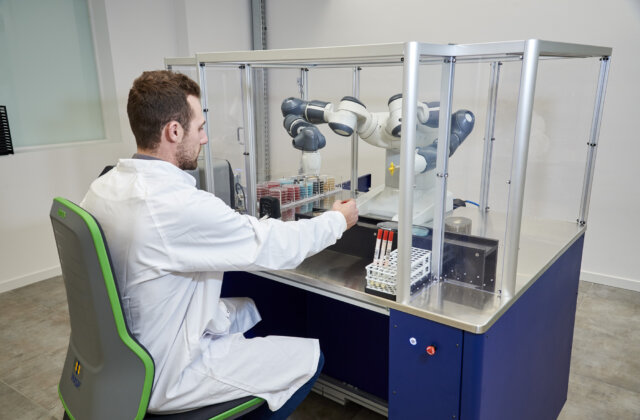
Colibrí™ is a fully automated instrument that prepares colonies for MALDI ID and susceptibility testing. When combined with TAG* functionality of PhenoMATRIX®, the software can analyze digital plate images, identify colonies of interest according to laboratory-defined parameters, and communicate colony coordinate information to the Colibrí™ to support automated ID and AST workflows.
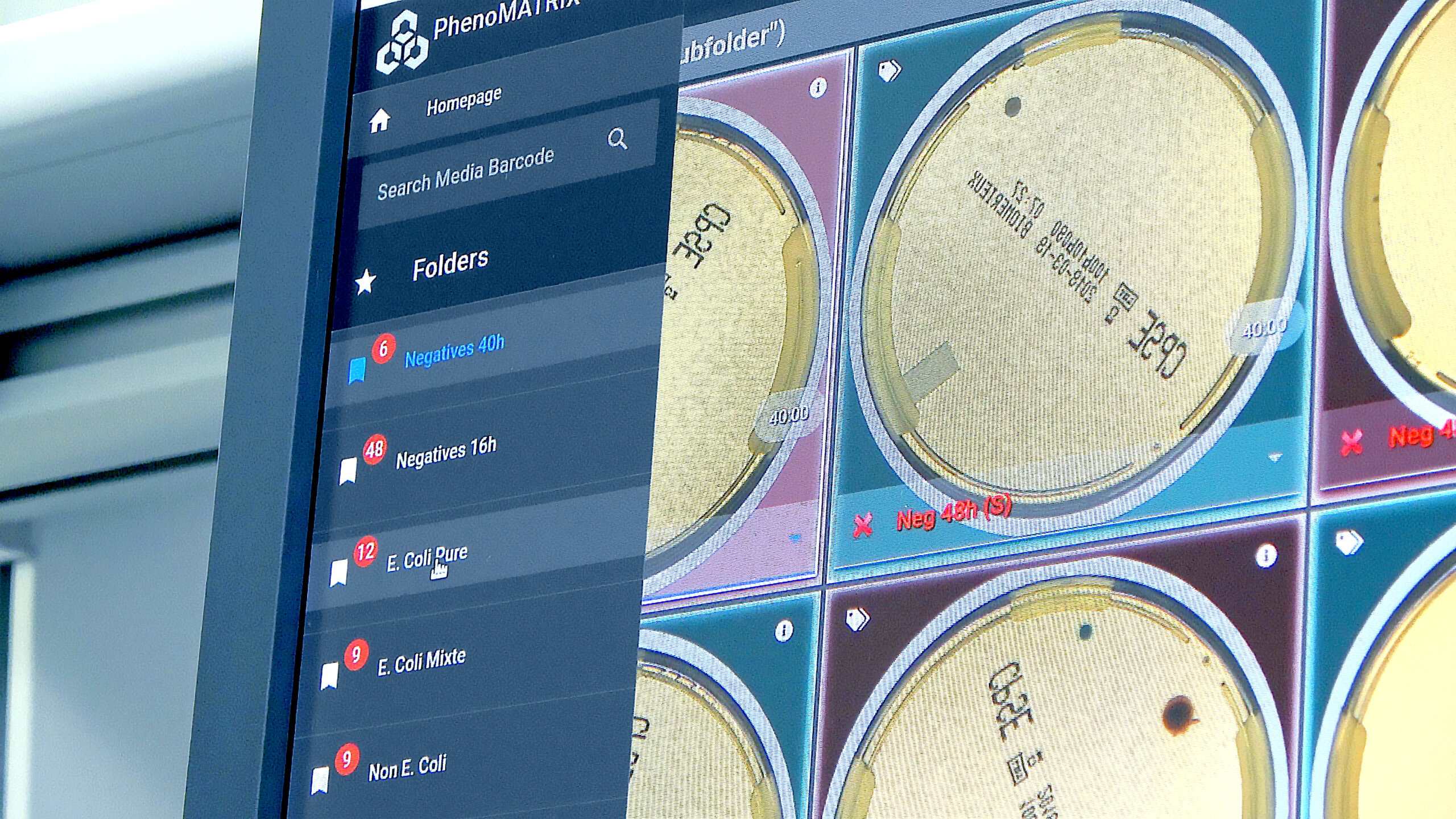
PhenoMATRIX® with Auto-Release
PLUS* functionality of PhenoMATRIX® is designed to streamline microbiology workflows by automatically sorting and releasing negative plates directly to the LIS without the need for manual review. The system flags plates with potential positives for technician validation while routing insignificant or negative results for automated release.**
PhenoMATRIX® Videos
Dynacare's Journey with Copan Automation & AI
In this illuminating video, you'll hear directly from the experts at DynaCare about embracing Copan's revolutionary artificial intelligence software – PhenoMATRIX®.
PhenoMATRIX® Laboratory Site Tour
PhenoMATRIX® and Digital Microbiology are Redefining What's Possible in Clinical Microbiology.
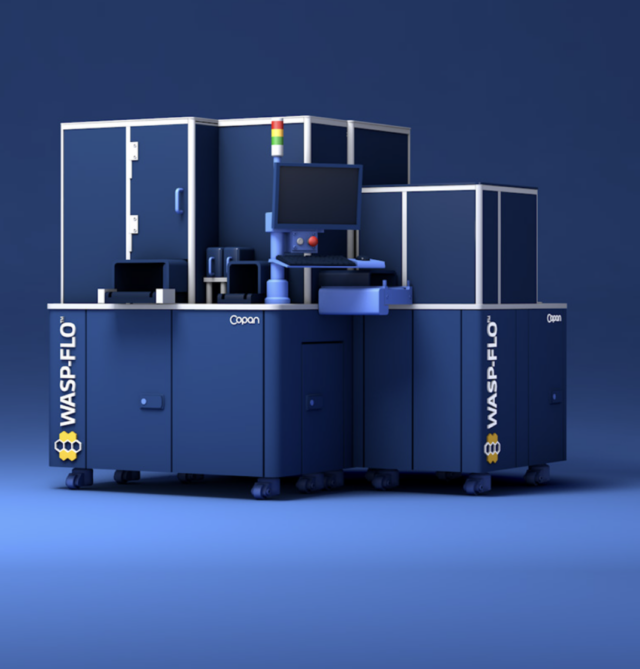
We help you begin your automation journey
Copan delivers full laboratory automation innovations spanning specimen sorting, processing, smart incubation, specialized imaging, and AI interpretation to maximize clinical microbiology productivity. Our modular platforms are uniquely customized to each laboratory while enhancing quality, safety and traceability.
References & Citations
1. Davidson RJ, et al. ASM Poster. 2024. AI for Identifying Urine Culture Results Automatically Released to Patient Records
2. Grams TR, et al. 2024. Evaluation of AI Software for Screening of Stool Cultures
3. Cherkaoui A, et al. J Clin Microbiol. 2023. Evaluation of PhenoMATRIX for MRSA Screening
4. Rovira-Pluja J, et al. ECCMID Poster. 2024. VITEK MS Prime Bacterial Identification Performance with Automated Slide Preparation
5. Rovira-Pluja J, et al. ECCMID Poster. 2024. How TLA and AI Could Improve Urine Culture Management
6. Dauwalder O, et al. Clin Microbiol Infect. 2021;27:1168.e1-1168.e6. Use of AI for Tailored Routine Urine Analyses
*Product labeled for Research Use Only (RUO) in the United States. Availability and regulatory status may vary by country.
**Laboratories remain responsible for establishing, validating, and overseeing any auto-verification or release policies in accordance with applicable regulations and laboratory protocols.
Resources and Downloads
PhenoMATRIX®
Resources and Downloads
Our Promise
Copan is committed to innovation in preanalytics. The automation developed by the COPAN team helps Microbiologists to provide faster and better results, and ultimately to make a difference and have a strong impact on patient care.