Scientific Studies
Thank you for visiting Copan’s Scientific Studies Library, your online resource for proof sources related to Copan products as well as many other aspects of sample collection and transport, automation, food and pharmaceutical testing, forensic and genetic collection and more!
Some of the studies in this database are independent scientific studies performed by third parties, while some of the studies were partially or entirely funded by Copan. In addition to peer reviewed research this library also contains posters, which were presented at various scientific meetings, and may not have been subject to peer review.
Some of the studies found here may contain information about off label product use. Studies, product claims and availability may vary from country to country, and they are subject to varying regulatory requirements. Always refer to product labeling, package inserts and safety data sheets for the appropriate use of the products.
Recent Scientific Studies
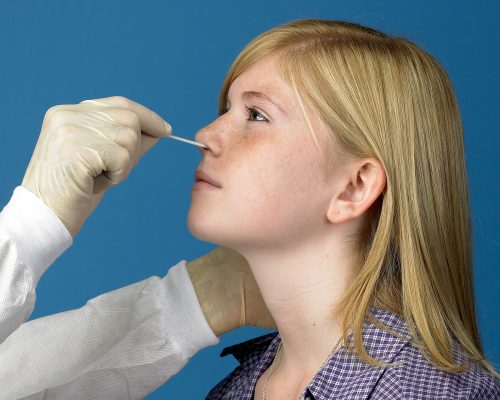
Recent Study Finds Nasopharyngeal (NP) Swabs Remain the Gold Standard for Diagnosis of COVID-19
April 28, 2021

Study Shows COPAN eNAT™ Transport System Can Address Challenges in COVID-19 Diagnostics in Regions with Limited Testing Access
February 15, 2021
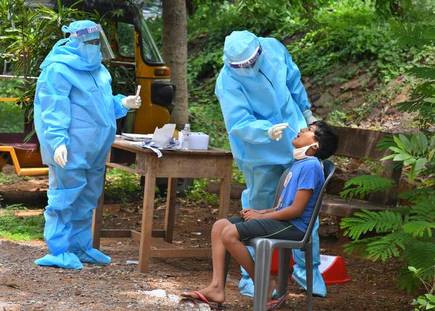
New Study Shows the Impacts of the Variability in Instructions for Performance of Nasopharyngeal Swabs in the Era of COVID-19
February 11, 2021